Fall of the Roman Empire, polygenic score edition
A first look at this puzzle from a genetic perspective
We have a new study out:
Piffer, D., Dutton, E., & Kirkegaard, E. O. W. (2023). Intelligence Trends in Ancient Rome: The Rise and Fall of Roman Polygenic Scores. OpenPsych, 1(1). https://doi.org/10.26775/OP.2023.07.21
We analysed 127 Ancient Roman genomes with a view to understanding the possible reasons for the fall of the Roman Empire. Taking the polygenic score for educational attainment (EA4) as a proxy for intelligence, we find that intelligence increased from the Neolithic Era (Z= -0.77) to the Iron Age (Z= 0.86), declines after the Republic Period and during the Imperial Period (Z= -0.27) and increases in Late Antiquity (Z= 0.25) and is approximately at the same level today (Z= 0.08). We show that this is congruent with a cyclical model of civilization based around intelligence, with the documented history of Rome, and also with patterns of immigration into Rome.
There is a lot of historical speculation on the rise and fall of the Roman empire. Pretty much everything has been invoked (including by me, lead poisoning and heat stress). One book from 1984 mentioned 200+ proposed reasons, which you can read a list of here. The list includes genetics indirectly, e.g. "degeneration", "decline of the Italic population" (replacement migration), and "degeneration of intellect" (could be internal dysgenic, or environmental poisoning, or cultural decline). With genetic prediction models of psychological traits (GWASs) and ancient genomic datasets, it is possible to examine the role of genetics, whether from replacement migration or some kind of negative selection for intelligence or other traits. I already wrote a summary post of one of these large-scale studies that looked at ancient genomics in general, but in this study we looked at Rome in particular. I proposed back in 2017 to look at this question with ancient genomic data and we now finally found some data for this.
The source of the genomic data is this study:
Antonio, M., Gao, Z., Moots, H., Lucci, M., Candilio, F., et al. (2019). Ancient rome: A genetic crossroads of europe and the mediterranean. Science, 366, 708–714.
Ancient Rome was the capital of an empire of ~70 million inhabitants, but little is known about the genetics of ancient Romans. Here we present 127 genomes from 29 archaeological sites in and around Rome, spanning the past 12,000 years. We observe two major prehistoric ancestry transitions: one with the introduction of farming and another prior to the Iron Age. By the founding of Rome, the genetic composition of the region approximated that of modern Mediterranean populations. During the Imperial period, Rome’s population received net immigration from the Near East, followed by an increase in genetic contributions from Europe. These ancestry shifts mirrored the geopolitical affiliations of Rome and were accompanied by marked interindividual diversity, reflecting gene flow from across the Mediterranean, Europe, and North Africa.
Here's their overview:
This glances over considerable complexity in where the bones originate from. One can look at their supplementary table for details:
So the 11 samples from the Roman Republic are actually from various places and subcultures.
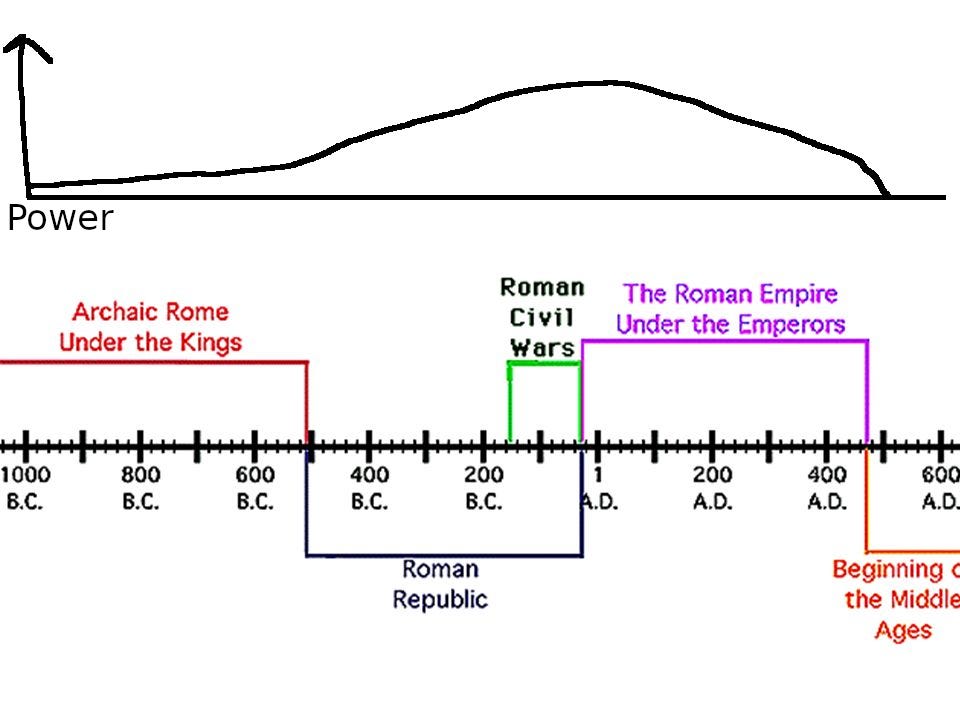
In terms of a genetic theory of the rise of fall of ancient Rome, one would expect something like this:
Where the power of Rome is a kind of extended phenotype of the average Roman genome's intelligence and other relevant traits. In fact, the effect of genes should be somewhat delayed because of the time it takes people to grow up and attain political power, and also because institutions are somewhat sticky. Ideally, one would plot a running mean of the average polygenic score for our best proxy for this, that of education (EA4). However, since the data for time is very course (carbon or cultural artifacts dating), we cannot do this for now, but we can at least try to approximate it by plotting the means by period:
The plot is of course noisy, there are only 127 genomes available. Nevertheless, the Republic is the highest average, as predicted by this theory. Imperial is low, whereas one would expect it to be somewhat elevated. This depends on whether the genomes are from the actual Romans or from recent immigrants (for whom one would expect a lower polygenic score). The difference between the Republic and the other periods is far beyond chance level (model p = .0002):
In the second model, the principal component 1 is controlled for. What does it mean?
As we explain in the paper:
A 2-component principal component analysis was run on the individual genomes. Three outliers at the far extreme of PC1 were identified and removed from the analysis. There was significant overlap between the time periods, although the second component clearly differentiated the pre-iron age period from the others (Figure 2).
The PC1 scores were lowest in the imperial period, and higher in the republican and mediaeval periods. Thus, they can be interpreted as Western European ancestry, which was lower during the imperial period but higher in the republican and mediaeval periods (Antonio et al., 2019). PC1 was significantly correlated to EA4 (r = 0.36, p= 3.5 × 10-5) but there was no correlation between PC2 and EA4 (r= 0.05, p= 0.558).
OLS linear regression was run to test the model that polygenic scores are significantly different between groups after accounting for population stratification (first principal component, PC1). A significant model emerged (R2= 0.18, p= 0.0002). PC1 was a significant predictor of the eduPGS (Beta= 0.28, p < 0.008), but among the periods, only the Republican period was a significant predictor (Beta= 0.96, p< 0.01) (Table 2).
So European ancestry (Italian in this case) predicts the EA4 score, but even controlling for this, the Republic period was still higher.
Various other comparisons, such as that between Republic vs. non-Republic (0.94 d), or Republic vs. modern (0.94 d) were also large in effect size and with small p values.
Conclusions
Based on existing ancient genomes from ancient Rome, there is support for education polygenic score differences between periods of the civilization. This follows predictions from a cyclical theory of selection for and against intelligence.
Evidence is not impressively strong owing to the small numbers of genomes (only 11 for Republic!), though p values were mostly reasonably small (below .01).
Same studies should be done for other great civilizations of the past for which a rise and fall in polygenic scores could be expected: ancient Greece, Persian empires, Egyptian empires, Mayans. It's an exciting time to come.
As a bonus, we can see that the right people are offended:
It is not exactly obvious why anti-science Adam Rutherford is unhappy. There's not much race science here, mainly, the theory is about the internal dynamics of Italians. We already know that modern Europeans were under positive and since around 1850-1900, are under negative selection for intelligence. What's the fuss about? My guess is that he's unhappy because (genetic) social cycle theories are somewhat inconsistent with a straight reading of history as one of endless progress.








If there are dysgenic or eugenic trends, then it is easy to see why populations must differ in average. Plus, he thinks that opens a door for eugenism. Those are the main reasons of Adam's unhappiness.
Reading your study synopsis, a thought comes to mind—what is meant by “immigrants”? I say this because in the last stages of the Republic, the influx of slaves from newly conquered Gaul and other provinces was huge. I’ve read as much as 40% of the population in Italy were slaves. Indeed, Octavian was much beloved because he passed a law requiring farmers to *hire* a percentage of Roman citizens as vs all slaves. Not sure how this changes findings, but it could sure account for a quick injection of foreign DNA into a population.