Genomic prediction of faces
Kickass paper by Xiong et al 2025
I sometimes get asked when we are going to get face accurate prediction from genomes. I tell people that it will take a while because variation is mostly continuous (unlike eye colors, which is very clustered) and there aren't many single variants of large effect size. I'm happy to say that there is new exciting progress in this area.
Xiong, Z., Li, Y., Liu, X., Lu, H., Hysi, P. G., Pardo, L. M., ... & Kayser, M. (2025). Combined genome-wide association study of facial traits in Europeans increases explained variance and improves prediction. Nature Communications, 16(1), 1-19.
Facial appearance, one of the most recognizable and heritable human traits, exhibits substantial variation across individuals within and between populations due to its complex genetic underpinning, which remains largely elusive. Here, we report a combined genome-wide association study (C-GWAS) of 946 facial features derived from 44 landmarks obtained from 3D digital facial images of 11,662 individuals of European descent. We identify 253 unlinked single nucleotide polymorphisms (SNPs) across 188 distinct genetic loci significantly associated with facial variation, including 64 SNPs at 62 novel loci and 33 novel SNPs within 29 previously reported face loci that are in very low LD with the previously reported top SNPs. Together, these SNPs account for up to 7.9% of the facial variation per trait, marking an average 2.25-fold increase over previous estimates. Cross-ancestry replication in 9,674 Chinese confirms the effect of 70% of these SNPs. A 382-SNPs prediction model of five nose traits achieves an AUC of 0.67 for individual re-identification from nose images. DNA predicted faces of archaic humans differ more from those of Europeans than from Africans. In genetically modelled Neanderthal faces, 15 of 16 DNA-predicted facial features are in line with skull evidence. Ten DNA-predicted facial features differentiate Neanderthals from Denisovans. Overall, this study substantially enhances our genetic understanding of human facial variation and provides improvements of genetic face prediction in modern and archaic humans.
A few things of note. First, it is amusing how the abstract just casually claims that face variation is heritable between populations (races) as the opening line (how do they know they aren't due to purely socioeconomic factors as well?). Second, the sample size is tiny compared to the usual GWASs, only 12k Europeans and an additional 10k Chinese.
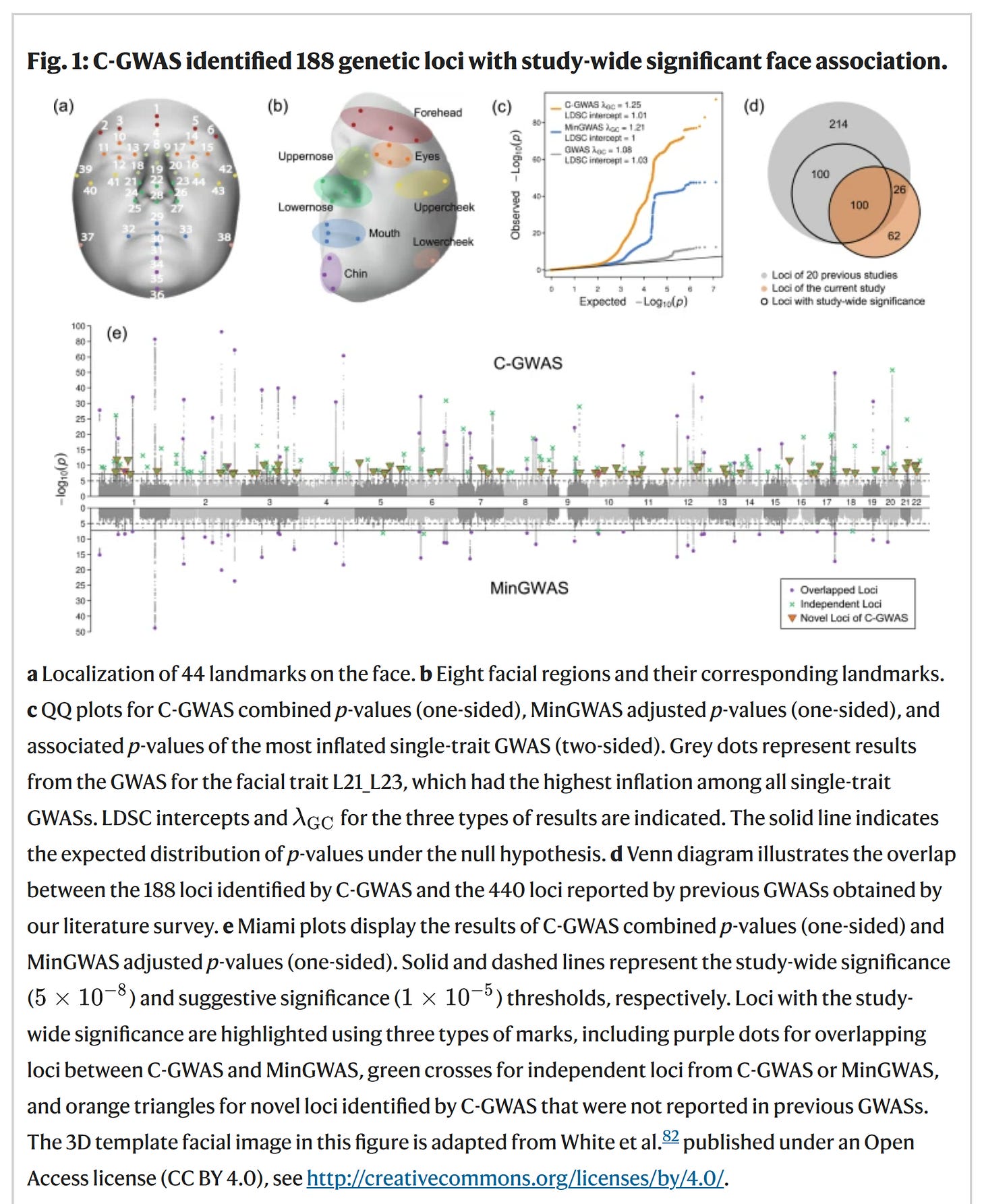
The way computers think about faces is not the way humans think about faces (at least, it doesn't appear that way). The first step is to analyze a bunch of faces and find the places they vary the most, so-called facial landmarks. Looks like this:
When the landmarks are identified, one can conduct the usual GWASs of each of them. Interesting to note was:
SNP-based heritability of 514 facial traits GWAS was estimated using LD score regression34 (LDSC) at an average of 0.23 and a range from 0.06 to 0.36 (Supplementary Data 2). Higher heritability was observed for the nose and forehead, whereas lower values for the mouth and cheek (Supplementary Fig. 2). Notably, the mouth and chin exhibited significant heritability divergence between internal variation and distances to other facial regions, highlighting the composite nature of facial variation, driven by cranial structure and soft tissue thickness35.
So even facial features show quite small snp heritabilities (average 23%). This underscores the usual worry that snp heritabilities dramatically underestimate overall heritabilities for whatever reasons (non-measured variants, poorly imputed variants, faulty methods, or faulty statistical assumptions), since of course no one thinks the real heritability of facial features is only 23% (identical twins look extremely similar to the point of getting confused for each other).
The variants and genes these implicated are also related to other phenotypes. Such genetic overlap (pleiotropy) can be the reason why people can predict psychological variation from facial features, that is, physiognomy:
Hierarchical clustering analysis revealed four distinct sets of candidate genes predominantly affecting different sets of facial regions (Fig. 3b and Supplementary Fig. 9) and showed specificity in biological processes (Fig. 3a and Supplementary Data 9). For example, genes influencing the nose were more enriched in limb morphogenesis, while those affecting the chin and mouth were more enriched in morphogenesis of branching structures, particularly axons. These findings offer a preliminary map of how embryonic development, driven by shared genetic factors, potentially influences distinct facial features. The extensive pleiotropy was further supported by a GWAS Catalog look-up, where 144 (57%) of the 253 lead SNPs, or their high LD counterparts (r2 > 0.6), were associated with 13 phenotypic categories of non-facial traits (Supplementary Fig. 10). The top-associated categories are anthropometric, brain imaging, metabolic and appearances traits, such as height, waist-to-hip ratio, sulcal depth, cortical surface area, total testosterone levels, contactin-2 levels, male pattern baldness, and hair colour (Supplementary Data 11). These results collectively suggest that the phenotypic spectrum of face-associated SNPs is more extensive than previously may have thought.
In terms of predictive validity at the individual level, we will still have to wait a while for something impressive:
Across the 514 facial traits, the combined proportion of variance explained (PVE) from all 253 lead SNPs together ranged from 2.25% to 7.89% (Supplementary Data 2), a 2.25-fold average increase from our earlier study3, which were based on 31 SNPs and 78 facial traits and ranged from 0.65% to 4.62%.
Recall that you need to use the square root of these values to interpret them properly. As such, in the best case scenario, a correlation of r = 0.28 was obtained between a genetic prediction and the landmark values among faces. This is not too bad, and is enough for group-level comparisons and even embryo selection. The authors elaborate:
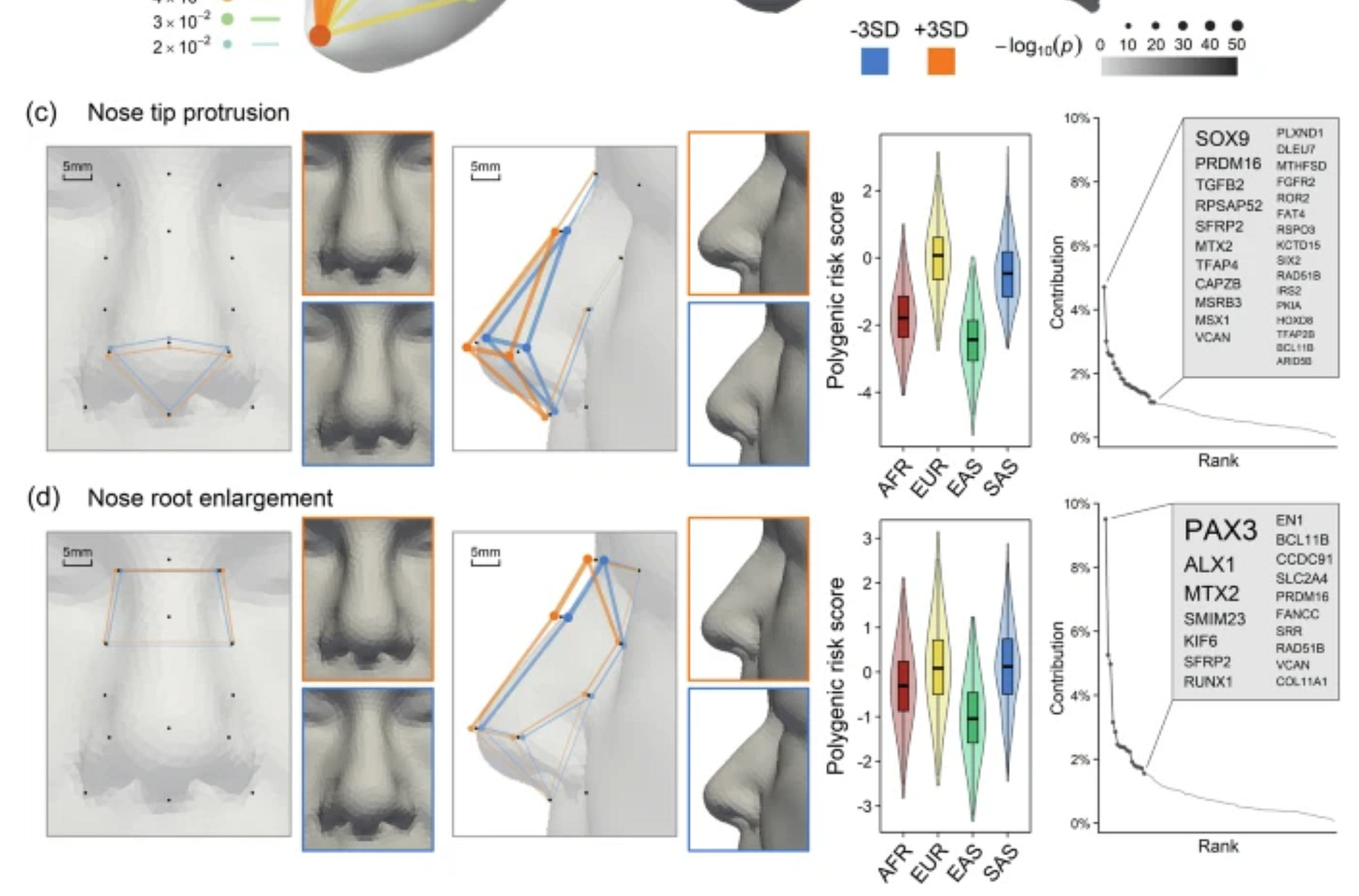
Building on these findings, we constructed polygenic risk score (PRS) profiles for 2000 individuals across four major continental populations (African, AFR; European, EUR; East Asian, EAS; South Asian, SAS) using data from the 1000 Genomes Project40, excluding admixed samples from AFR and the American population. The PRS was based on 382 face-associated SNPs, including 253 lead SNPs from our C-GWAS and 129 SNPs from previous facial shape GWASs, for 23 most genetically explainable and independent facial traits (see the “Methods” section, Supplementary Data 13). Facial PRS profiles largely agree with established anthropological knowledge concerning nose shape variation among continental populations (Fig. 4b–d, Supplementary Data 14 and Supplementary Note 6). A comparative analysis of computed tomography scans of 388 adults of AFR, Asian (ASN), and EUR ancestry41 further confirmed significant correlations between mean differences in facial PRS profiles and phenotypic traits between EUR and AFR (Pearson’s r = 0.75), as well as between EUR and ASN (r = 0.81) (Supplementary Note 7). Specifically, considering the average facial PRS profiles of EUR as reference (z = 0), AFR showed notably smaller nose root (z = −1.86), less protruded (z = −1.76) but more upturned (z = 0.74) nose tips, alongside broader nose wings (z = 0.8) and shorter nose bridges (z = −0.82). EAS were characterized by significantly smaller (z = −1.77) and less (z = −2.42) protruded nose tips, along with smaller nose root (z = −1.12). Meanwhile, SAS displayed similar nose shapes to EUR (all |z | <0.5), which is remarkable given their closer geographic proximity to EAS. The latter finding is in line with the study of Zaidi et al.42 finding that the average nose differences between SAS and EUR were smaller than other intercontinental comparisons. Furthermore, our findings are largely consistent with numerous facial photogrammetric studies reviewed by Wen et al.5, reinforcing the correlation between genetic facial profiles and physical facial anthropometry.
Visually:
The correlations at the group level (group means correlations) are particularly important to heredidtarians as they indirectly confirm our results using a new independent test conducted by mainstream researchers working with unrelated phenotypes. Supplementary note 7 reads:
To evaluate whether our PRS capture facial shape differences across modern human populations, we conducted a comparative analysis using 3D facial data. We focused on the study by SimmonsEhrhardt et al.14, which analysed 3D bone and soft tissue parameters from computed tomography scans of 388 adults from AFR, Asian (ASN), EUR, and Hispanic populations. This study was selected because it used 3D data, as we used in our study, and included many facial traits that overlapped with those in our dataset. We identified 42 facial phenotypes common to both studies. For each trait, we calculated the standardized mean differences between non-European populations (AFR, and ASN, excluding Hispanic) and the EUR, using the EUR as the reference. This resulted in 42×2 phenotype mean differences representing the facial shape variations among populations. We constructed PRS for the same 42 traits using 382 SNPs and the same approach described in Methods “Genetic modelling of facial traits”. These PRS were applied to individuals from corresponding populations in the 1000 Genomes Project dataset. We calculated the standardized mean differences in PRS between non-European and European populations, yielding 42×2 PRS mean differences. Note that because the study of Simmons-Ehrhardt et al. does not distinguish EAS and SAS, we merged in our study EAS and SAS as ASN samples.
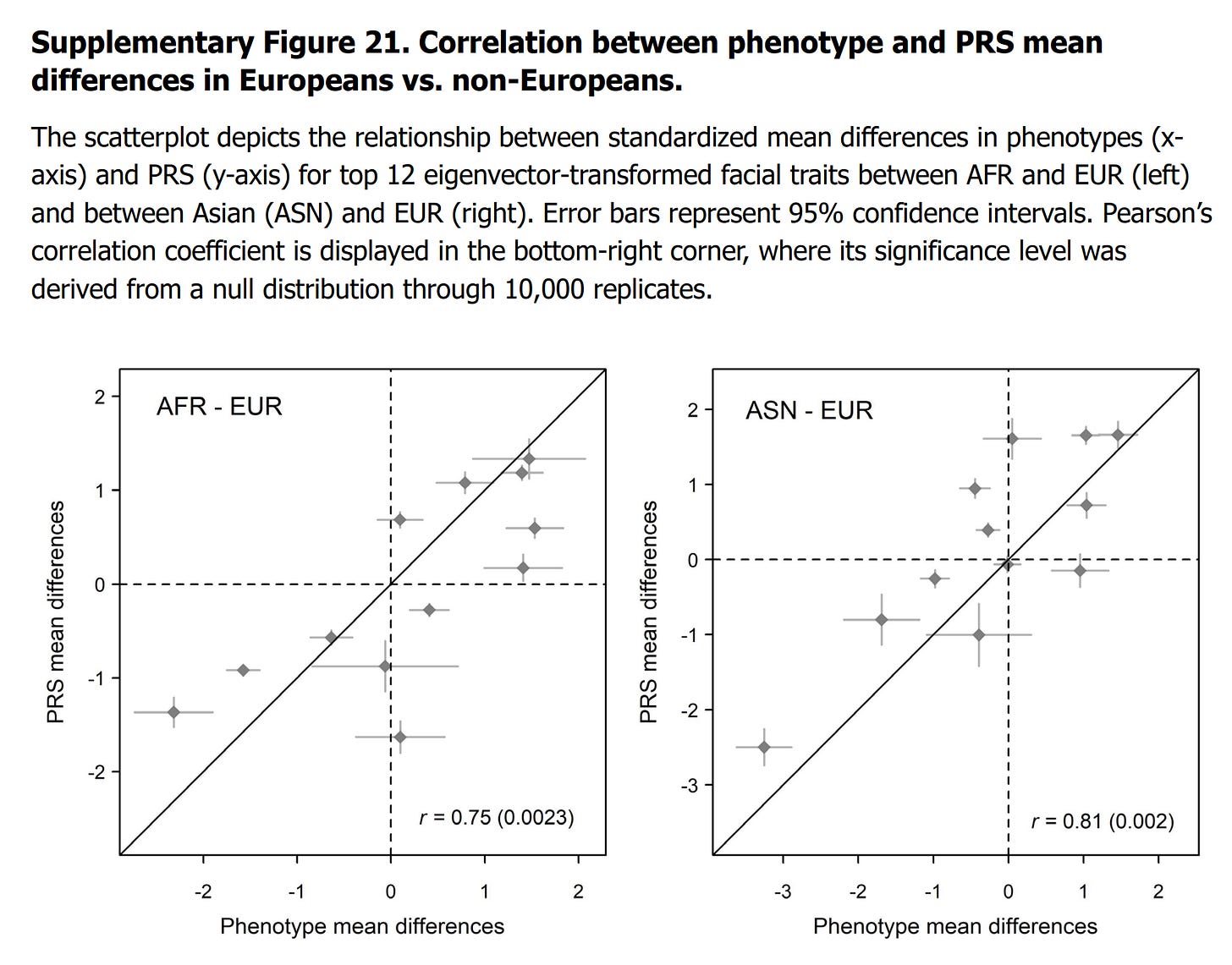
Since many of the 42 traits are highly correlated, we transformed the PRS and phenotype mean differences into an eigenvector space using the same approach described in Methods “Proportion of Facial Variance Genetically Explained”. We retained the top 12 transformed mean differences, corresponding to the accumulated Eigen value over 95% of total 42. We assessed the correlation between the 12 transformed PRS mean differences and the phenotype mean differences. A higher correlation would indicate that the PRS effectively reflects the observed facial differences among populations. To determine the significance of this correlation, we generated a null distribution through 10,000 replicates. In each replicate, we constructed PRS using 382 randomly selected SNPs that were matched for MAF. We transformed mean differences and calculated the correlations for these replicates to establish a null distribution of correlations expected by chance. We compared our observed correlation to the null distribution to calculate a p-value. This allowed us to assess whether the PRS captured population-level facial differences beyond what would be expected randomly. The analysis revealed significant correlations between mean differences in facial traits and mean differences in PRS between EUR and AFR (Pearson’s r = 0.75, p = 0.0023), as well as between EUR and ASN (r = 0.81, p = 0.002) populations (Supplementary Fig. 21).
Thus, they used the same statistical testing approach as we regularly use (random MAF-matched snps). This approach is necessary because of the correlated nature of the snps and human populations are genetically related, resulting in genetic autocorrelation. This means true null correlations at the group level will have a much wider distribution (spread around 0) than under regular statistical assumptions, and thus correlations need to be very strong to beat the null. Nevertheless, they report good correlations of 0.75 and 0.81, which look like this:
Unfortunately, they didn't label the points on the plots.
To be noted here, the dataset they had is the reverse structure of what we normally work with. Normally, we have only a few phenotypes+PGS combinations (EA, IQ, height, BMI) but many samples (worldwide, or subnational regions). They did the opposite, many phenotype+PGS pairs and only pairs of groups (3 groups total). So their analysis was whether the polygenic scores correctly identify the correct differences among two pairs of populations (Europeans vs. African, Europeans vs. Asians) across many phenotypes, whereas we regularly work with testing whether the polygenic scores correctly rank order many different populations or subpopulations on a single phenotype. The latter method is more challenging because the use of many populations make worries about population stratification worse.
Next up, they checked whether these facial variants have been under selection. Europeans certainly differ more among each other in coloration traits than other races, especially north Europeans (blue eyes, red hair etc.). These populations are also the ones the highest in individualism (Hajnal-line/WEIRD) mentality. This could be a coincidence, but it is probably not. Individualism as a cultural trait is more useful the easier it is to facially distinguish people, and this ability can be enhanced if there is more genetic facial variation:
Among the 253 lead SNPs identified in our facial C-GWAS, 69% exhibited a frequency difference > 0.2 among EUR, EAS, and AFR, as per the 1000 Genomes Project data (Supplementary Data 4). This proportion is significantly higher than the average observed in random samples of the same number of SNPs across the genome after 10,000 replicates (p = 0.023, Supplementary Fig. 12). The average of fixation index (FST), which reflects genetic variation between groups and can indicate local positive selection, was significantly higher between EUR and EAS (p = 0.017) and EUR and AFR (p = 0.011) than expected from randomly sampled SNPs (Fig. 6a). However, this pattern was not observed in the EAS-AFR comparison (p = 0.32, Supplementary Fig. 12). Moreover, the mean Population Branch Statistic (PBS) for the 253 lead SNPs compared to randomly selected SNPs was statistically significant in EUR (p < 0.01, Fig. 6a), but not in EAS (p = 0.44) and AFR (p = 0.25). These findings suggest a more pronounced influence of positive selection on face-associated SNPs in Europeans.
Some of the p-values aren't that great, but there appears to be some signal in this direction. One worry here is that this GWAS was carried out in Europeans, thus inflating the MAF of the variants in the models (statistical power depends on the frequencies of the snps under study; it is hard to find a very small p value for a snp that is very rare to begin with). However, against this worry, the authors note that:
Although our targeted SNPs may have higher MAF in EUR because they were identified through a European-based C-GWAS, a recent large-scale face GWAS conducted in East Asians22 identified 244 face-associated SNPs and found evidence of positive selection on these SNPs in Europeans. This suggests that facial differences between Europeans and East Asians, such as more protruded and narrow noses in Europeans, may result from adaptation and positive selection in European populations. The consistency of findings across independent GWAS in different continental populations indicates these results are not merely artefacts of GWAS-based allele frequency biases.
I also note that people working in facial recognition have told me that their models have higher accuracies for Europeans.
What about ancient humans? The authors looked at this as well:
Next, we constructed facial PRS profiles based on 357 face-associated SNPs available in 10 archaic humans (8 Neanderthals, 1 Denisovan, and 1 Neanderthal–Denisovan admixed individual) using genomic data obtained from the Allen Ancient DNA Resource54 (see the “Methods” section, Supplementary Table 3 and Supplementary Data 18). When comparing archaic humans to the four major modern human populations in the 1000 Genomes Project, archaic facial PRS profiles are more different from three non-African groups, including EUR, than from AFR (Fig. 7c). This finding aligns with current understanding that Africans are ancestral to all modern humans, making them genetically closer to the common ancestors of both modern and archaic humans. However, this finding may appear as a paradox given that the introgression happened in non-Africans. To explore this further, we calculated the mean absolute differences in allele frequencies for the 357 SNPs between archaic humans and modern human populations, and compared these to 10,000 random sets of number-matched random SNPs. Differences between archaic humans and AFR were consistently smaller than those between archaic humans and all non-Africans from EUR, EAS, and SAS, even when focusing on SNPs within introgressed segments (Fig. 7d). These findings suggest that the similarity in facial PRS profiles between archaic humans and AFR is likely due to shared ancestral alleles. However, we acknowledge a potential bias stemming from the predominantly European GWAS origin of these 357 facial SNPs. Future studies should prioritize facial GWAS in diverse populations, particularly in African cohorts, to reduce such biases and clarify the evolutionary history underlying facial shape genetics.
Somewhat surprisingly, they find that Africans are more similar to Neanderthals/Denisovans than Europeans and Asians are (East and South). This is despite the fact that the non-African populations have a small amount of genetic admixture from these ancient populations (2-4%). However, from the perspective of statistical genetics, it is not so surprising since Africans are closer genetically to Neanderthals/Denisovans than non-Africans are (all the groups split off from the original African population but at different times).
Since we also have Neanderthal skulls, we can check whether the predictions work, and they do:
Neanderthal skulls have been discovered and show a series of distinct features that are different compared to skulls of modern humans51,55. To validate Neanderthal facial PRS profiles, we mapped them to 16 Neanderthal facial features reflected by skulls, excluding features not reflected in bone structure, such as the nose tip and certain aspects of the nose bridge (Supplementary Table 4, see the “Methods” section). Of the 16 facial features analysed, 15 showed concordances between (a) the direction of the difference predicted by facial PRS profiles in Neanderthals relative to Europeans, and (b) the direction of the reported feature differences for Neanderthals relative to modern humans, examples include a wider face, more protruded brow ridge, flatter cheekbones, and lower palate (Fig. 8a). Permutation tests (see the “Methods” section) indicated that observing 15 concordant features is statistically highly significant, exceeding all 10,000 random permutations (p < 0.0001, Fig. 8b). Only one feature (wider palate in Neanderthals) showed inconsistent classification due to conflicting directions from multiple overlapping PRS (p = 0.0126). No discordant results were observed (p = 0.0012). The statistical significance of these findings indicates that the observed concordance between European-based PRS predictions for Neanderthals and their known facial features relative to modern humans is unlikely to have occurred by chance. To further assess the robustness of these results, we conducted simulations to evaluate how data limitations for archaic humans, such as pseudohaploid data, small sample sizes, and low call rates, might affect PRS accuracy (Supplementary Notes 10 and 11). The results showed that pseudohaploid data and small sample sizes did not bias PRS averages, although they did increase the uncertainty. Additionally, imputing genotypes for low call rate samples yielded PRS averages highly consistent with those from high call rate samples. These findings suggest that, despite moderate PRS r2 values that may not accurately represent individual facial profile and the inherent uncertainty of archaic DNA data, the broader patterns in PRS profiles, such as directionality of averages, remained robust, particularly for extreme values, reinforcing the reliability of archaic PRS profiles for characterizing population-level facial difference.
They also draw the same inference as we generally do: you don't need a lot of predictive validity in PGSs to correctly find the group differences.
All in all, this is a great study. A good candidate for one of the most important genomics studies this year so far.


Can you publish a statement on Erik Ahrens? This guy is now apparently sharing your info as well as his other contacts with Antifa and suddenly claims to have renounced his far-right beliefs to support open borders and multiculturalism and says he no longer opposes mass immigration but wants to destroy Germany by a melting-pot. Now calls openly himself a "feminist", promotes OnlyFans and has a LGBT flag on his X profile. Is this another case of horseshoe theory or an elaborate trolling/parody? Or was he influenced by Hanania? He does not appear to have yet abandoned HBD and appears to be combining HBD with now far-left/progressive politics.
Terrible article. Filled with errors. Do better, emil.